













| General Information: |
|
| Name(s) found: |
MSB1_YEAST
[Swiss-Prot]
|
| Description(s) found: |
|
| Organism: | Saccharomyces cerevisiae S288c |
| Length: | 1137 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MNDMAKPLPT PPTAEIRKSR SNSPKKAQKT NLSPNKNQNN EKNVPRSNGR TKNEHNSMDD 60
61 EEFEFFHQFS REKVKGVIHV ITAELKEKGP DVEFLMIPFR PEQTNDKLLT LLNQLFPLGN 120
121 GQPVNEKKQL RIVSKADVWT LFQCLKYIWC RLPNSEIIGW KSYLEFKFRE EDKKFPRKSF 180
181 LEIMPQCLAS PNHASIVYDF FDLIISISSN SRVNKMSARK ISKMCAIWAF SKQIPNSDIQ 240
241 DYDFESAAMK SFAPNNSIQD GLDQWIPASD AMFHLLLAFL RSFVPQDLES AKLPRTLKSL 300
301 LFNNQYPPRK STAYTSETIL TIPLVTLKTD VFSRKPWQLL ERCNDLLDFS DHDAFEARED 360
361 YALLKSLFRK KNTVEGISRK MSQESRRLMK AMSTKHSTFQ PGWAPRECIE NISHLKECIE 420
421 VKRLDIDDYF IWTWLSSLSF EQTSEKKKIF GRSIILEFEF DGFKKWVVFQ ECDITLDYNK 480
481 KGQLKKKTSA QSPTTEKELP PDDFELEDPP LSKSPTLSQT YKKFQAEVPQ QSTVRRDSAP 540
541 DNQGIYHTVI SKNALTKNKH NVNLHSFEHK ISKWNPLNNL RKKSGSNSSS SSFEEKSKDA 600
601 PIREEYHTNK NHKSKKEERV LSQFSTLNPD EYQLPVIETG SSNFKIEIPE LMYEHDDDDS 660
661 DKLKNSQKRA TDSAIEELNG MVEEMMINEP DDVKISITEA ETFESLTKFD QYKPSNITDD 720
721 DLQSSHSSAV HSLKLSTNTN DSCADSSKYT ADRKLAEPRK ISEESKVNDD SSSYYSPNIN 780
781 NLPASRMPSQ PTYSNSDSKK AFTNESRLNV LQGAVSPSQQ VTPKPYKNAP GDCVSPVQQK 840
841 YYQNDRRNEM SPASAPVPPS AYSPARSPQF STNSAGFKQN TINVPVGYND PAHVLANQPH 900
901 MTYRDQHNYP SHQQKQRPFQ NNIVPPELKS RNQRADASPI PQHMVPVKQG VPNLPSNVPL 960
961 YQQMERMNPN HQHPVNTYKV TQPPYHNNTT NAYGNSRAGN AHMLDGKWSN NPPQMVPKGV1020
1021 RPNQYPQQHV NRYSPQAQPV VPAEYYNGPP PMRAPPMMSH MVPAQEPIRY TAGANRRSFP1080
1081 QGMQQNAYSV PAQPMGAVNS EFYLPEAPQG NKLHGNINKR QERKKLYDNI RSGNFGI |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
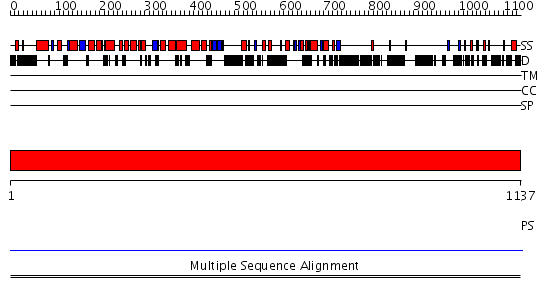
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..1137] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [309..588] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [589..771] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [772..929] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [930..1137] | N/A | No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||
| 5 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)