













| General Information: |
|
| Name(s) found: |
ENO2_ARATH
[Swiss-Prot]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Arabidopsis thaliana |
| Length: | 444 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA]
chloroplast [IDA] cytoplasm [IDA] membrane [IDA] apoplast [IDA] mitochondrion [IDA] mitochondrial envelope [IDA] plasma membrane [IDA] |
| Biological Process: |
response to cold
[IMP]
response to abscisic acid stimulus [IEP] response to salt stress [IEP] response to cadmium ion [IEP] response to light stimulus [IMP] |
| Molecular Function: |
phosphopyruvate hydratase activity
[IDA][ISS]
copper ion binding [IDA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MATITVVKAR QIFDSRGNPT VEVDIHTSNG IKVTAAVPSG ASTGIYEALE LRDGGSDYLG 60
61 KGVSKAVGNV NNIIGPALIG KDPTQQTAID NFMVHELDGT QNEWGWCKQK LGANAILAVS 120
121 LAVCKAGAVV SGIPLYKHIA NLAGNPKIVL PVPAFNVING GSHAGNKLAM QEFMILPVGA 180
181 ASFKEAMKMG VEVYHHLKSV IKKKYGQDAT NVGDEGGFAP NIQENKEGLE LLKTAIEKAG 240
241 YTGKVVIGMD VAASEFYSED KTYDLNFKEE NNNGSQKISG DALKDLYKSF VAEYPIVSIE 300
301 DPFDQDDWEH YAKMTTECGT EVQIVGDDLL VTNPKRVAKA IAEKSCNALL LKVNQIGSVT 360
361 ESIEAVKMSK KAGWGVMTSH RSGETEDTFI ADLAVGLSTG QIKTGAPCRS ERLAKYNQLL 420
421 RIEEELGSEA IYAGVNFRKP VEPY |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
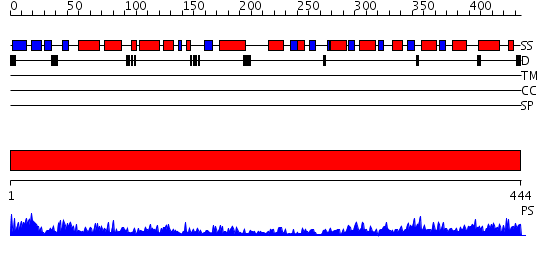
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..444] | 158.0 | Crystal Structure of Human Neuron Specific Enolase at 1.8 angstrom |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)