













| General Information: |
|
| Name(s) found: |
DED1 /
YOR204W
[SGD]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 604 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IC |
| Biological Process: |
RNA splicing
[IDA translational initiation [IMP |
| Molecular Function: |
RNA helicase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAELSEQVQN LSINDNNENG YVPPHLRGKP RSARNNSSNY NNNNGGYNGG RGGGSFFSNN 60
61 RRGGYGNGGF FGGNNGGSRS NGRSGGRWID GKHVPAPRNE KAEIAIFGVP EDPNFQSSGI 120
121 NFDNYDDIPV DASGKDVPEP ITEFTSPPLD GLLLENIKLA RFTKPTPVQK YSVPIVANGR 180
181 DLMACAQTGS GKTGGFLFPV LSESFKTGPS PQPESQGSFY QRKAYPTAVI MAPTRELATQ 240
241 IFDEAKKFTY RSWVKACVVY GGSPIGNQLR EIERGCDLLV ATPGRLNDLL ERGKISLANV 300
301 KYLVLDEADR MLDMGFEPQI RHIVEDCDMT PVGERQTLMF SATFPADIQH LARDFLSDYI 360
361 FLSVGRVGST SENITQKVLY VENQDKKSAL LDLLSASTDG LTLIFVETKR MADQLTDFLI 420
421 MQNFRATAIH GDRTQSERER ALAAFRSGAA TLLVATAVAA RGLDIPNVTH VINYDLPSDV 480
481 DDYVHRIGRT GRAGNTGLAT AFFNSENSNI VKGLHEILTE ANQEVPSFLK DAMMSAPGSR 540
541 SNSRRGGFGR NNNRDYRKAG GASAGGWGSS RSRDNSFRGG SGWGSDSKSS GWGNSGGSNN 600
601 SSWW |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | McCusker D, et al (2007) | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | De Craene, J., et al. (2006) | |
| View Run | No Comments | Widlund PO, et al. (2005) | |
| View Run | No Comments | Widlund PO, et al. (2005) | |
| View Run | his-HA tag on RPA135 | Schneider, DA, et al. (2006) | |
| View Run | his-HA tag on RPA135 | Schneider, DA, et al. (2006) | |
| View Run | Dam1-765 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | control 1 | McCusker D, et al (2007) | |
| View Run | control 2 | McCusker D, et al (2007) | |
| View Run | experimental1 - mih1-1 | McCusker D, et al (2007) | |
| View Run | experimental2 - mih1-2 | McCusker D, et al (2007) | |
| View Run | cac1: TAP-tagged | Green EM, et al (2005) | |
| View Run | Dam1-765 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | phosphorylation data - cascade search | Green EM, et al (2005) | |
| View Run | identification data - first search in cascade | Green EM, et al (2005) | |
| View Run | sample 3928 - Asf1-S-TEV-ZZ (in hir3delta mutant) | Green EM, et al (2005) | |
| View Run | Sample 2- cpn2 from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample 3 - cpn + atp, from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample boi1 with gst from october 2005 | McCusker D, et al (2007) | |
| View Run | Sample bob1 (2nd set) from october 2005 | McCusker D, et al (2007) | |
| View Run | Boi 2 gst control | McCusker D, et al (2007) | |
| View Run | Sample: HX11 - both fusion and trans-SNARE complex formation take place. | Hao Xu, et al. (2010) | |
| View Run | Sample: HX12 - trans-SNARE complex formation and fusion are inhibited (control). | Hao Xu, et al. (2010) | |
| View Run | Sample: HX14 - mixture in detergent (control). | Hao Xu, et al. (2010) | |
| View Run | #25 Asynchronous Prep3-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #03 Alpha Factor Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #13 Mitotic Prep2-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #12 Mitotic Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #10 Mitotic Prep1-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #02 Alpha Factor Prep1-TiO2 enriched, new search criteria | Keck JM, et al. (2011) | |
| View Run | #30 Asynchronous Prep6-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #11 Mitotic Prep1-TiO2 enriched, new search criteria | Keck JM, et al. (2011) | |
| View Run | #06 Alpha Factor Prep3-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #08 Alpha Factor Prep4-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
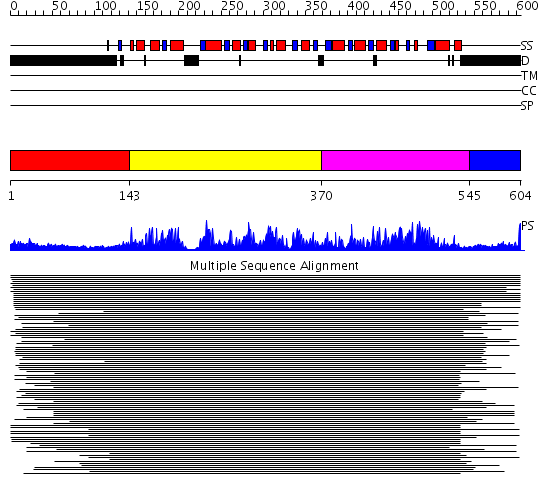
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..142] | 7.22 | Fibrinogen | |
| 2 | View Details | [143..369] | 672.218487 | Putative DEAD box RNA helicase | |
| 3 | View Details | [370..544] | 672.218487 | Putative DEAD box RNA helicase | |
| 4 | View Details | [545..604] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.94 |
Source: Reynolds et al. (2008)