













| General Information: |
|
| Name(s) found: |
SPR1 /
YOR190W
[SGD]
|
| Description(s) found:
Found 29 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 445 amino acids |
Gene Ontology: |
|
| Cellular Component: |
fungal-type cell wall
[IMP |
| Biological Process: |
ascospore formation
[IMP |
| Molecular Function: |
glucan 1,3-beta-glucosidase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MVSFRGLTTL TLLFTKLVNC NPVSTKNRDS IQFIYKEKDS IYSAINNQAI NEKIHGVNLG 60
61 GWLVLEPYIT PSLFETFRTN PYNDDGIPVD EYHFCEKLGY EKAKERLYSH WSTFYKEEDF 120
121 AKIASQGFNL VRIPIGYWAF TTLSHDPYVT AEQEYFLDRA IDWARKYGLK VWIDLHGAAG 180
181 SQNGFDNSGL RDSYKFLEDE NLSATMKALT YILSKYSTDV YLDTVIGIEL LNEPLGPVID 240
241 MERLKNLLLK PAYDYLRNKI NSNQIIVIHD AFQPYHYWDG FLNDEKNEYG VIIDHHHYQV 300
301 FSQVELTRKM NERIKIACQW GKDAVSEKHW SVAGEFSAAL TDCTKWLNGV GLGARYDGSW 360
361 TKDNEKSHYI NTCANNENIA LWPEERKQNT RKFIEAQLDA FEMTGGWIMW CYKTENSIEW 420
421 DVEKLIQLNI FPQPINDRKY PNQCH |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Qiu et al. (2008) |

|
CRR1 SPR1 |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
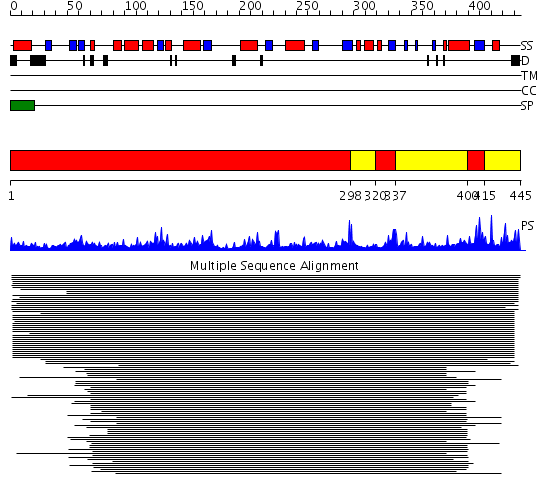
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..297] [320..336] [400..414] |
1328.69897 | Exo-beta-(1,3)-glucanase | |
| 2 | View Details | [298..319] [337..399] [415..445] |
1328.69897 | Exo-beta-(1,3)-glucanase |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Source: Reynolds et al. 2008. Manuscript submitted |

|