













| General Information: |
|
| Name(s) found: |
SHM2 /
YLR058C
[SGD]
|
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 469 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[TAS |
| Biological Process: |
one-carbon metabolic process
[TAS |
| Molecular Function: |
glycine hydroxymethyltransferase activity
[TAS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MPYTLSDAHH KLITSHLVDT DPEVDSIIKD EIERQKHSID LIASENFTST SVFDALGTPL 60
61 SNKYSEGYPG ARYYGGNEHI DRMEILCQQR ALKAFHVTPD KWGVNVQTLS GSPANLQVYQ 120
121 AIMKPHERLM GLYLPDGGHL SHGYATENRK ISAVSTYFES FPYRVNPETG IIDYDTLEKN 180
181 AILYRPKVLV AGTSAYCRLI DYKRMREIAD KCGAYLMVDM AHISGLIAAG VIPSPFEYAD 240
241 IVTTTTHKSL RGPRGAMIFF RRGVRSINPK TGKEVLYDLE NPINFSVFPG HQGGPHNHTI 300
301 AALATALKQA ATPEFKEYQT QVLKNAKALE SEFKNLGYRL VSNGTDSHMV LVSLREKGVD 360
361 GARVEYICEK INIALNKNSI PGDKSALVPG GVRIGAPAMT TRGMGEEDFH RIVQYINKAV 420
421 EFAQQVQQSL PKDACRLKDF KAKVDEGSDV LNTWKKEIYD WAGEYPLAV |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Krogan NJ, et al. (2006) |

|
SHM2 YEL047C |
| View Details | Gavin AC, et al. (2002) |
|
GCN20 GFA1 SHM2 |
| View Details | Ho Y, et al. (2002) |
|
ADK1 CLU1 COP1 GPH1 NAN1 PRP11 REX2 RFC3 RFC4 RFC5 RPN11 RPT3 SEC27 SHM2 SSK2 TFP1 THI22 TIF4631 TMA29 UBP15 UTP13 YCL042W YGR250C |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | sample: hx1 | Hao Xu, et al. (2010) | |
| View Run | sample: hx2 | Hao Xu, et al. (2010) | |
| View Run | sample: hx3 | Hao Xu, et al. (2010) | |
| View Run | Sample: HX11 - both fusion and trans-SNARE complex formation take place. | Hao Xu, et al. (2010) | |
| View Run | Sample: HX12 - trans-SNARE complex formation and fusion are inhibited (control). | Hao Xu, et al. (2010) | |
| View Run | Sample: HX13 - trans-SNARE complex forms but fusion is blocked. | Hao Xu, et al. (2010) | |
| View Run | Sample: HX14 - mixture in detergent (control). | Hao Xu, et al. (2010) | |
| View Run | #24 Asynchronous Prep3-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #12 Mitotic Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
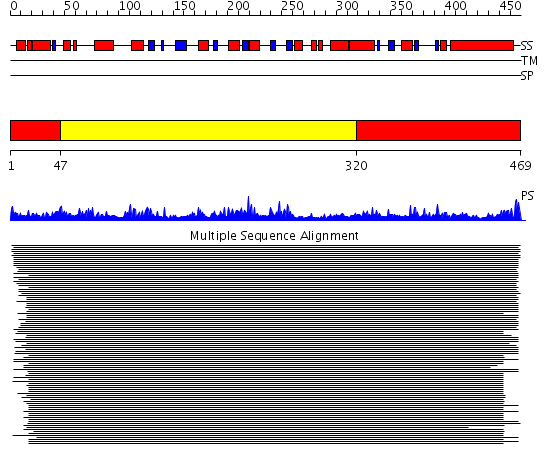
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..46] [320..469] |
1689.0 | Serine hydroxymethyltransferase | |
| 2 | View Details | [47..319] | 1689.0 | Serine hydroxymethyltransferase |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.89 |
Source: Reynolds et al. (2008)