













| General Information: |
|
| Name(s) found: |
UGA3 /
YDL170W
[SGD]
|
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 528 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IC |
| Biological Process: |
positive regulation of transcription, DNA-dependent
[IDA nitrogen utilization [IMP regulation of transcription from RNA polymerase II promoter [IMP |
| Molecular Function: |
transcription factor activity
[IDA specific RNA polymerase II transcription factor activity [IPI |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MNYGVEKLKL KYSKHGCITC KIRKKRCSED KPVCRDCRRL SFPCIYISES VDKQSLKKIK 60
61 ADIQHQLISK KRKHAPDSAQ KAAVATRTRR VGSDEQDNQV YLSKPLEDCI SQKLDSMGLQ 120
121 LYNYYRSHLA NIISIAPMNQ NYYLNIFLPM AHENDGILFA ILAWSANHLS ISSSNELRKD 180
181 EIFVNLANKY TYMSLSHLKT NEGSSACAKL GFLYSLAQIL ILCGSEICQG DVKFWKILLN 240
241 IGKNLIENHV GKDVSRILTT TTEEPSLEER IIFPNFNSVV KYWLIVNFIY HDILNFNTTS 300
301 FPIEQYEKFF QRDQNSLPSS ANFIESIDSP IEEIDPLIGI NKPILLLLGQ VTNLTRFLQT 360
361 MEQEEMLEHG DKILSLQVEI YKLQPSLMAL EHLDDEKKFY YLELFEIMKI STLMFFQLTL 420
421 LKIDKDSLEL QILRNKLDSK LDKVIGTFLE GSLCFPLFIY GVCIQVEDME KKIDLEAKFD 480
481 DILKRYKCYN FQNARLLIRK IWQNEADGIS EHDLVHMIDE LDYNINFA |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | sample cbf3 from feb 2005 | Sandall S, et all (2006) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
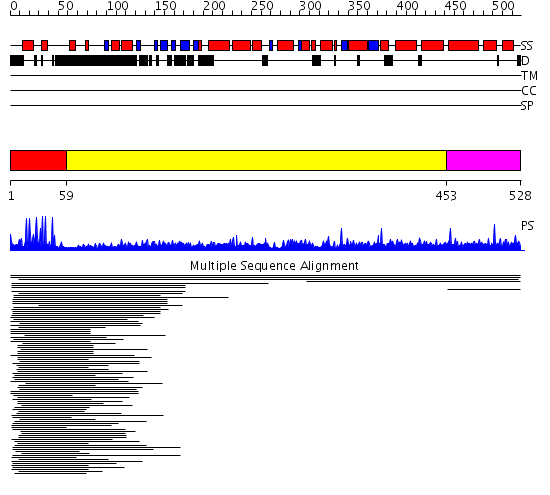
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..58] | 32.201249 | PPR1 | |
| 2 | View Details | [59..452] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [453..528] | 1.004969 | View MSA. Confident ab initio structure predictions are available. | |
| 4 | View Details | [395..528] | 1.032967 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||
| 3 | No functions predicted. | ||||||||||||||||||||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)