













| General Information: |
|
| Name(s) found: |
RNE_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 6 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 1061 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MKRMLINATQ QEELRVALVD GQRLYDLDIE SPGHEQKKAN IYKGKITRIE PSLEAAFVDY 60
61 GAERHGFLPL KEIAREYFPA NYSAHGRPNI KDVLREGQEV IVQIDKEERG NKGAALTTFI 120
121 SLAGSYLVLM PNNPRAGGIS RRIEGDDRTE LKEALASLEL PEGMGLIVRT AGVGKSAEAL 180
181 QWDLSFRLKH WEAIKKAAES RPAPFLIHQE SNVIVRAFRD YLRQDIGEIL IDNPKVLELA 240
241 RQHIAALGRP DFSSKIKLYT GEIPLFSHYQ IESQIESAFQ REVRLPSGGS IVIDSTEALT 300
301 AIDINSARAT RGGDIEETAF NTNLEAADEI ARQLRLRDLG GLIVIDFIDM TPVRHQRAVE 360
361 NRLREAVRQD RARIQISHIS RFGLLEMSRQ RLSPSLGESS HHVCPRCSGT GTVRDNESLS 420
421 LSILRLIEEE ALKENTQEVH AIVPVPIASY LLNEKRSAVN AIETRQDGVR CVIVPNDQME 480
481 TPHYHVLRVR KGEETPTLSY MLPKLHEEAM ALPSEEEFAE RKRPEQPALA TFAMPDVPPA 540
541 PTPAEPAAPV VAPAPKAAPA TPAAPAQPGL LSRFFGALKA LFSGGEETKP TEQPAPKAEA 600
601 KPERQQDRRK PRQNNRRDRN ERRDTRSERT EGSDNREENR RNRRQAQQQT AETRESRQQA 660
661 EVTEKARTAD EQQAPRRERS RRRNDDKRQA QQEAKALNVE EQSVQETEQE ERVRPVQPRR 720
721 KQRQLNQKVR YEQSVAEEAV VAPVVEETVA AEPIVQEAPA PRTELVKVPL PVVAQTAPEQ 780
781 QEENNADNRD NGGMPRRSRR SPRHLRVSGQ RRRRYRDERY PTQSPMPLTV ACASPELASG 840
841 KVWIRYPIVR PQDVQVEEQR EQEEVHVQPM VTEVPVAAAI EPVVSAPVVE EVAGVVEAPV 900
901 QVAEPQPEVV ETTHPEVIAA AVTEQPQVIT ESDVAVAQEV AEQAEPVVEP QEETADIEEV 960
961 VETAEVVVAE PEVVAQPAAP VVAEVAAEVE TVAAVEPEVT VEHNHATAPM TRAPAPEYVP1020
1021 EAPRHSDWQR PTFAFEGKGA AGGHTATHHA SAAPARPQPV E |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
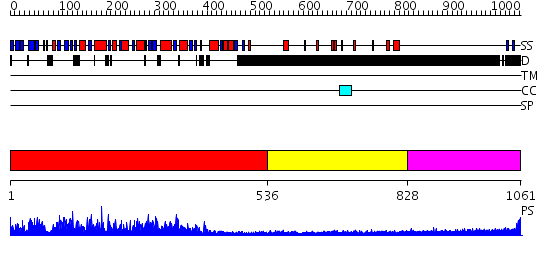
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..535] | 1000.0 | Catalytic domain of E. coli RNase E | |
| 2 | View Details | [536..827] | 2.09 | 30S subunit | |
| 3 | View Details | [828..1061] | 6.374997 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.91 |
Source: Reynolds et al. (2008)