













| General Information: |
|
| Name(s) found: |
gi|30725364
[NCBI NR]
gi|26450663 [NCBI NR] gi|25344801 [NCBI NR] gi|15222191 [NCBI NR] gi|10092282 [NCBI NR] |
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 322 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cellular_component
[ND]
|
| Biological Process: |
metabolic process
[IEA]
regulation of nitrogen utilization [IEA] |
| Molecular Function: |
oxidoreductase activity, acting on NADH or NADPH
[ISS]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTTEKSKILV IGGTGYMGEF IVEGSAKAGN PTFALVREAS LSDPVKSKTI QSFKDLGVTI 60
61 LHGDLNDHES LVKAIKQVDV VISTIGHKQI FDQTKIISAI KEAGNVKRFL PAEFGIDVER 120
121 TSAVEPAKSL FAGKVQIRRA IEAEGIPYTY VVSNCSAGFY LRTLLQFESG LISHTRDKAI 180
181 IFGDKNVPPR DKVTILGDGN AKVVINKEED VAAYMIKAVD DLRTLNKTLY ISPPNNILSM 240
241 NEMVTLWEKK IGKSLEKTHI SEEQILKSIQ VPIDVFKSIN HAVFVKGDQT SFTIEPWFGE 300
301 EASVLYPDVK YTSIDEYLSQ FT |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
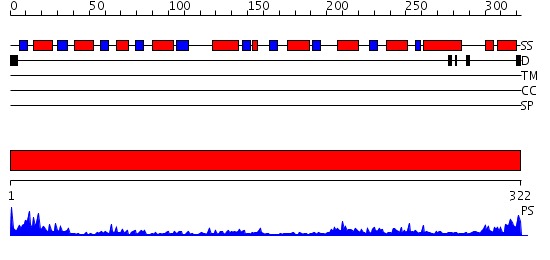
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..322] | 41.522879 | Crystal structures of pinoresinol-lariciresinol and phenylcoumaran benzylic ether reductases, and their relationship to isoflavone reductases |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.95 |
Source: Reynolds et al. (2008)