













| General Information: |
|
| Name(s) found: |
CE34135
[WormBase]
|
| Description(s) found:
Found 14 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 715 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQNFGGPGGP MYGGAGGGGG GPPRGPPQQP PQPQGGGVAV PFPIPQGITQ AHYHQYKQEL 60
61 TNFQQNRLAL LNQALNRGQL THEQFQQEVH NVNERIRGAM TQYAAQCSQR AAAAQQQQQQ 120
121 QNQAAGQMPP PQMAPVMMSA AEKKKQMEMQ KHQRMAQQQQ MQQQQGGMSF PPGSTLQQQQ 180
181 QQMRMQFQRQ QQQQQQQLQQ NPQNQGYGRM GGGPQYPGMQ QQQQLTHQNP GMQQQQMQQP 240
241 QMQQTPQSHG PIPVSNPSSA QQMQYYQQQQ QQQLQLQQNQ QMQMQQQQMQ QHQQQHPGMG 300
301 GPEMGGHMHA QQGAPGSQAA PVQQQQQIQQ QAPPQAPAEK TEDMKYKELL KEMKLQYLEA 360
361 LQGMQRRQAQ QKGLVQMVNI LEGDRIVSYD HLLSLKTPLH RLMTRDCPTF PLMEEIRKVV 420
421 FKKKEDREKA MLMSFVGEKD NLSAQVESAD KRLSEQSKDD DPMGVKPWRS VKHLTIRVPD 480
481 FVRNLTGNED RKAFLKRPRA VSAEEDEAVV GAKKVKEEEQ DDTSSQASED EDISLIQSEF 540
541 VIKTVFDSRK PWIVPPKARE ELSNISHWNV DEHCLPSCSA TPFIIVCIKS PSLLVNPLRI 600
601 SLPRTYPIDP ATVQFDRTFP QDSEHGPVLQ KLFEKNLGAK TTVRTMTDFV EAYCSACEEY 660
661 QIFMPKYNNE SSPEQSSQPG SAHSSQQQKT PEMAYHSRLP AAVSNPIIAQ KGIVI |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp1 IP from december2004 | Cheeseman, I.M., et al. (2005) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
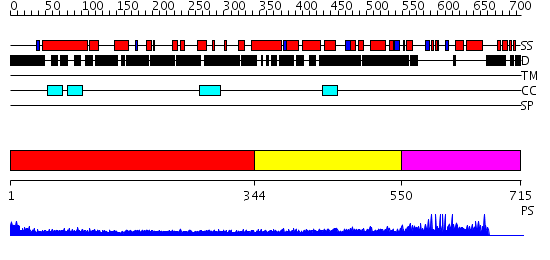
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..343] | 15.522879 | Heavy meromyosin subfragment | |
| 2 | View Details | [344..549] | 6.69897 | Tropomyosin | |
| 3 | View Details | [550..715] | 1.02 | Toxoplasma gondii ubiquitin conjugating enzyme TgTwinScan_2721- E2 domain |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||
| 1 | No functions predicted. | ||||||
| 2 | No functions predicted. | ||||||
| 3 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)