













| General Information: |
|
| Name(s) found: |
ntl-3 /
CE33892
[WormBase]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Caenorhabditis elegans |
| Length: | 701 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IEA |
| Biological Process: |
embryonic development ending in birth or egg hatching
[IMP regulation of transcription [IEA |
| Molecular Function: |
transcription regulator activity
[IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAEKRKLLAE IDKCFKKIDE GVELFEETME KMHEANSDNQ RDKYQDDLKK EIKKLQRLRD 60
61 QVKNWQNASE IKDKDKLNSY RKLIEQRMEQ FKDVERENKT KPHSKLGLSA EEKLDPKEKE 120
121 KAETMDWIQH QIRSLNEEVD RTEMQLESLS NTDTGKGKRG KKEDAKTKNE REKRVEGLKH 180
181 HLERINFHIE KLEICMRMIS NESLNAKMVL ETLKEPIETY VEMMNEEDSE EADNYDPDDA 240
241 YDELNLEKLC QQIGGVNVAS VDDEHRENGH ELGIDTAESG AVSGSRHTSG ENGQPPSPAG 300
301 RRIVPLSMPS PHAVTPELKR LASKDSNVDR PRTPPVTPAS AAPPPPGIPY NSVAAGRSTT 360
361 TPVPSTPISA NSPAPSLAQA APIAAASPVF PPAAAAASKP VLAQSVSEMP QKKESITSTT 420
421 SRGSAAAPAT TTTTTTTTTS SEPAEVPLVV QQTVSETFVN GVDSPAAATR LTQQERQQQL 480
481 QQQHHHQSTI IPTTPTTTTT SSSMLGGMMS TDDPAAALQA ALNMAAASQQ AVATGPKRAH 540
541 IPAWLGASPL GRTSMTQEFD GQLAALELAC AKATFPLDSE KPRNYLSKVS FPVPSWYGQT 600
601 APNTSDSLEY YLRLAPDTLF FIFYYMEGTR AQLLAAKALK KLSWRFHTKY LTWFQRHEEP 660
661 KQITDDYEQG TYVYFDFEKW SQRKKESFTF EYKFLEDKEF D |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | hcp2 ip from december | Cheeseman, I.M., et al. (2005) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
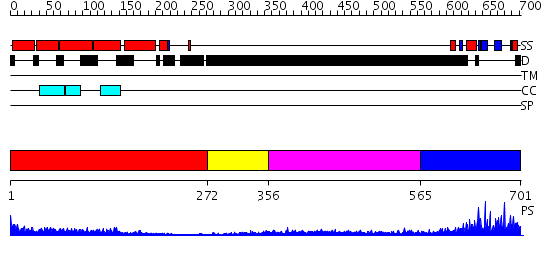
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..271] | 11.522879 | Tropomyosin | |
| 2 | View Details | [272..355] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [356..564] | 3.300997 | View MSA. No confident structure predictions are available. | |
| 4 | View Details | [565..701] | 30.853872 | No description for PF04153.9 was found. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)