













| General Information: |
|
| Name(s) found: |
UBP14_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 13 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 797 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cellular_component
[ND]
|
| Biological Process: |
protein deubiquitination
[IDA]
embryonic development ending in seed dormancy [IMP] |
| Molecular Function: |
ubiquitin-specific protease activity
[IDA][ISS]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MELLRSNLSR VQIPEPTHRI YKHECCISFD TPRSEGGLFV DMNSFLAFGK DYVSWNYEKT 60
61 GNPVYLHIKQ TRKSIPEDRP LKKPTLLAIG VDGGFDNNEP EYEESYSIVI LPDFVSLPFP 120
121 SVELPEKVRI AVDTVVNAVG AERKEQVAAW TAEKKLISEH ALTLQQIKSG IVIPPSGWKC 180
181 SKCDKTENLW LNLTDGMILC GRKNWDGTGG NNHAVEHYKE TAYPLAVKLG TITADLEAAD 240
241 VYSYPEDDSV LDPLLAEHLA HFGIDFSSMQ KTEMTTAERE LDQNTNFDWN RIQESGKELV 300
301 PVFGPGYTGL VNLGNSCYLA ATMQIVFSTH SFISRYFSHQ SLKMAFEMAP ADPTLDLNMQ 360
361 LTKLGHGLLS GKYSMPATQK DATTGDPRQE GIPPRMFKNV IAASHAEFSS MRQQDALDFF 420
421 LHLVGKVERA SNTTPDLDPS RSFKFGIEEK ILCPSGKVGY NKREDCILSL NIPLHEATNK 480
481 DELEAFHKQK AGKGLEENDM RSSDEIVRPR VPLEACLANF ASSEPIEDYY SSALKGMTTA 540
541 IKTTGLTSFP DYLVLHMRKF VMEEGWVPKK LDVYIDVPDV IDISHMRSKG LQPGEELLPD 600
601 GVPEEVMESA QPVANEEIVA QLVSMGFSQL HCQKAAINTS NAGVEEAMNW LLSHMDDPDI 660
661 DAPISHQTSD IDQSSVDTLL SFGFAEDVAR KALKASGGDI EKATDWVFNN PNASVSDMDV 720
721 SSSNSAQTPA QSGLPDGGGK YKLFGIVSHM GTSVHCGHYV AHILKEGRWV IFNDDKVGIS 780
781 TDPPKDMGYV YFFQRLD |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
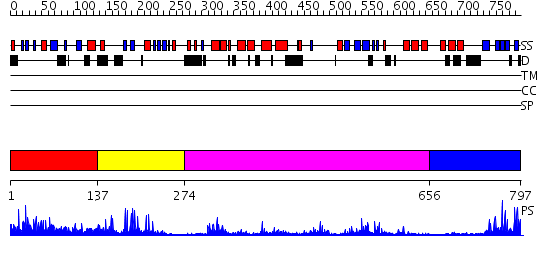
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..136] | 1.4 | No description for 2g43A was found. | |
| 2 | View Details | [137..273] | 24.09691 | No description for 2g43A was found. | |
| 3 | View Details | [274..655] | 49.0 | No description for 2gfoA was found. | |
| 4 | View Details | [656..797] | 4.89 | Crystal structure of HAUSP |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)